Provides estimation and plotting tools for exposure-response models that use logistic regression for binary responses. It is mostly intended as a convenience package: the core tools are wrappers around glm(), and the plotting tools use ggplot2 and patchwork to build typical plots used in exposure-response modelling.
Models
library(erlr)
library(tibble)
lr_data
#> # A tibble: 300 × 10
#> id sex age weight dose treatment aucss cmaxss ae1 ae2
#> <int> <fct> <int> <dbl> <dbl> <fct> <dbl> <dbl> <dbl> <dbl>
#> 1 1 Male 35 79 200 Drug 673. 97.3 0 1
#> 2 2 Female 22 58 200 Drug 2806. 301. 1 1
#> 3 3 Female 28 58 0 Placebo 0 0 0 0
#> 4 4 Female 18 57 100 Drug 1169. 198. 1 1
#> 5 5 Male 28 77 100 Drug 377. 51.4 0 0
#> 6 6 Female 19 76 200 Drug 327. 25.4 1 0
#> 7 7 Male 30 70 0 Placebo 0 0 0 0
#> 8 8 Female 34 60 100 Drug 1208. 133. 1 1
#> 9 9 Male 21 89 0 Placebo 0 0 0 0
#> 10 10 Female 34 56 200 Drug 254. 31.0 0 0
#> # ℹ 290 more rows
mod <- lr_model(ae1 ~ aucss, lr_data)
mod
#>
#> Call: stats::glm(formula = formula, family = stats::binomial(link = "logit"),
#> data = data)
#>
#> Coefficients:
#> (Intercept) aucss
#> -1.791383 0.005497
#>
#> Degrees of Freedom: 299 Total (i.e. Null); 298 Residual
#> Null Deviance: 402.1
#> Residual Deviance: 193.4 AIC: 197.4Plots
lr_data |>
lr_plot(aucss, ae1) |>
lr_plot_show_model() |>
lr_plot_show_quantiles() |>
lr_plot_show_groups(aucss) |>
plot()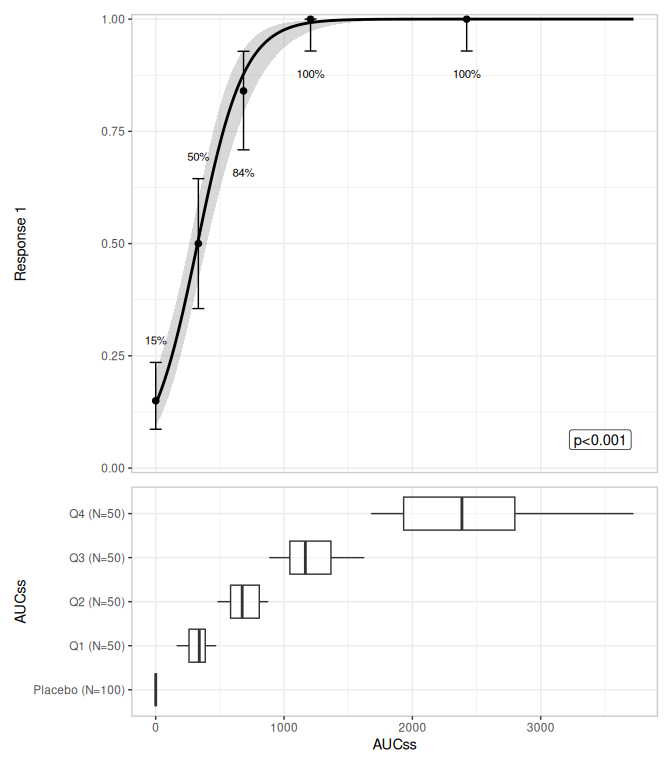
plt <- lr_data |>
lr_plot(aucss, ae2, stratify_by = sex) |>
lr_plot_show_model(keep_strata = FALSE) |>
lr_plot_show_quantiles(bins = 3) |>
lr_plot_show_datastrip() |>
lr_plot_show_groups(group_by = c(aucss, treatment), keep_strata = FALSE)
print(plt)
#> <erlr_plot>
#> plot variables:
#> - exposure: aucss
#> - response: ae2
#> - stratification: sex
#> plot components:
#> - model: ae2 ~ aucss
#> - quantile: 3 bins
#> - strip: jitter both
#> - group: .aucss_quantile, treatment
#> plots built: <none>
#> output built: no
plot(plt)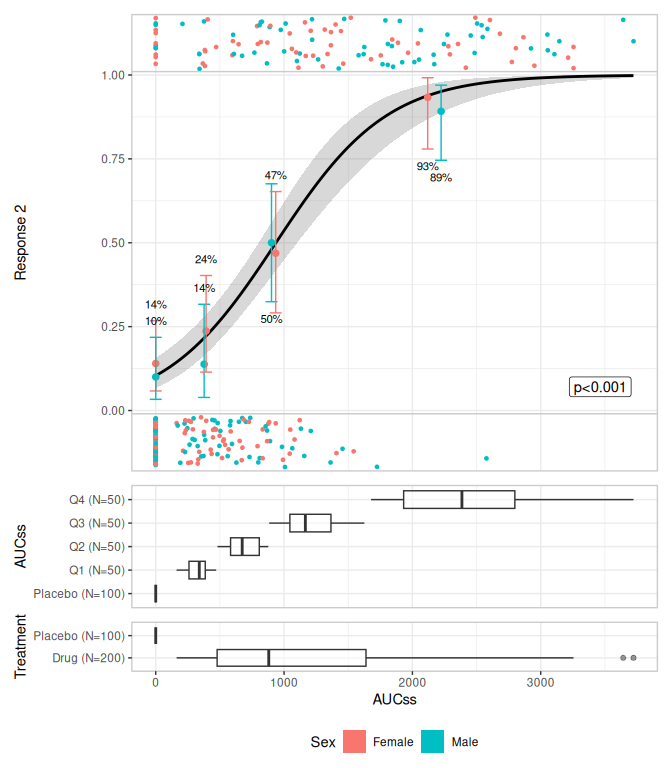
Stepwise covariate modelling
mod1 <- lr_model(ae1 ~ aucss + sex + dose, lr_data)
mod2 <- lr_scm_backward(mod1, candidates = c("sex", "dose"))
#> Using seed = 7932
lr_scm_history(mod2)
#> # A tibble: 4 × 11
#> iteration attempt step action term_tested model_tested model_converged
#> <int> <int> <chr> <chr> <chr> <chr> <lgl>
#> 1 0 0 base model <NA> <NA> ae1 ~ aucss +… TRUE
#> 2 1 1 backward remove ~dose ae1 ~ aucss +… TRUE
#> 3 1 2 backward remove ~sex ae1 ~ aucss +… TRUE
#> 4 2 3 backward remove ~sex ae1 ~ aucss TRUE
#> # ℹ 4 more variables: term_p_value <dbl>, model_aic <dbl>, model_bic <dbl>,
#> # model_updated <int>VPC/Simulation
mod <- lr_model(ae1 ~ aucss + sex, lr_data)
sim <- lr_vpc_sim(mod, seed = 1234)
sim
#> # A tibble: 30,000 × 5
#> ae1 aucss sex row_id sim_id
#> <dbl> <dbl> <fct> <int> <int>
#> 1 0.894 673. Male 1 1
#> 2 1.00 2806. Female 2 1
#> 3 0.110 0 Female 3 1
#> 4 0.993 1169. Female 4 1
#> 5 0.588 377. Male 5 1
#> 6 0.468 327. Female 6 1
#> 7 0.129 0 Male 7 1
#> 8 0.994 1208. Female 8 1
#> 9 0.129 0 Male 9 1
#> 10 0.362 254. Female 10 1
#> # ℹ 29,990 more rows
lr_vpc_plot(mod, sim, group_by = aucss)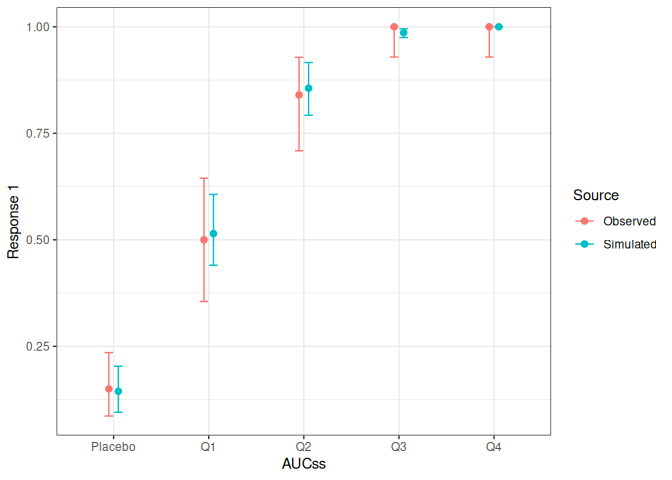
lr_vpc_plot(mod, sim, group_by = sex)